diff --git a/README.MD b/README.MD
index b76dfc6..d4c7691 100644
--- a/README.MD
+++ b/README.MD
@@ -1,5 +1,6 @@
[](https://github.com/Samyssmile/edux/actions/workflows/gradle.yml)
[](https://github.com/Samyssmile/edux/actions/workflows/codeql-analysis.yml)
+
# EDUX - Java Machine Learning Library
EDUX is a user-friendly library for solving problems with a machine learning approach.
@@ -8,18 +9,22 @@ EDUX is a user-friendly library for solving problems with a machine learning app
EDUX supports a variety of machine learning algorithms including:
-- **Multilayer Perceptron (Neural Network):** Suitable for regression and classification problems, MLPs can approximate non-linear functions.
+- **Multilayer Perceptron (Neural Network):** Suitable for regression and classification problems, MLPs can approximate
+ non-linear functions.
- **K Nearest Neighbors:** A simple, instance-based learning algorithm used for classification and regression.
- **Decision Tree:** Offers visual and explicitly laid out decision making based on input features.
- **Support Vector Machine:** Effective for binary classification, and can be adapted for multi-class problems.
- **RandomForest:** An ensemble method providing high accuracy through building multiple decision trees.
### Battle Royale - Which algorithm is the best?
+
We run all algorithms on the same dataset and compare the results.
[Benchmark](https://github.com/Samyssmile/edux/discussions/42)
## Goal
-The main goal of this project is to create a user-friendly library for solving problems using a machine learning approach. The library is designed to be easy to use, enabling the solution of problems with just a few lines of code.
+
+The main goal of this project is to create a user-friendly library for solving problems using a machine learning
+approach. The library is designed to be easy to use, enabling the solution of problems with just a few lines of code.
## Features
@@ -31,43 +36,122 @@ The library currently supports:
- Support Vector Machine
- RandomForest
-## Installation
+## Get started
Include the library as a dependency in your Java project file.
### Gradle
```
- implementation 'io.github.samyssmile:edux:1.0.5'
+ implementation 'io.github.samyssmile:edux:1.0.6'
```
### Maven
+
```
io.github.samyssmileedux
- 1.0.5
+ 1.0.6
```
-## How to use this library
+## Getting started tutorial
+
+This section guides you through using EDUX to process your dataset, configure a multilayer perceptron (Multilayer Neural
+Network), perform training and evaluation.
+
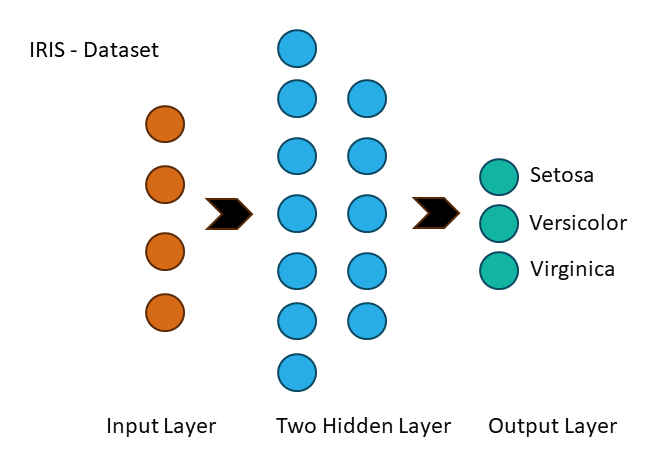
+A multi-layer perceptron (MLP) is a feedforward artificial neural network that generates a set of outputs from a set of
+input features. An MLP is characterized by several layers of input nodes connected as a directed graph between the input
+and output layers.
+
+
+
+### Step 1: Data Processing
+
+Firstly, we will load and prepare the IRIS dataset:
+
+| sepal.length | sepal.width | petal.length | petal.width | variety |
+|--------------|-------------|--------------|-------------|---------|
+| 5.1 | 3.5 | 1.4 | 0.2 | Setosa |
+
+```java
+var featureColumnIndices=new int[]{0,1,2,3}; // Specify your feature columns
+ var targetColumnIndex=4; // Specify your target column
+
+ var dataProcessor=new DataProcessor(new CSVIDataReader());
+ var dataset=dataProcessor.loadDataSetFromCSV(
+ "path/to/your/data.csv", // Replace with your CSV file path
+ ',', // CSV delimiter
+ true, // Whether to skip the header
+ featureColumnIndices,
+ targetColumnIndex
+ );
+ dataset.shuffle();
+ dataset.normalize();
+ dataProcessor.split(0.8); // Replace with your train-test split ratio
+```
+
+### Step 2: Preparing Training and Test Sets:
- NetworkConfiguration networkConfiguration = new NetworkConfiguration(...ActivationFunction.LEAKY_RELU, ActivationFunction.SOFTMAX, LossFunction.CATEGORICAL_CROSS_ENTROPY, Initialization.XAVIER, Initialization.XAVIER);
- MultilayerPerceptron multilayerPerceptron = new MultilayerPerceptron(features, labels, testFeatures, testLabels, networkConfiguration);
- multilayerPerceptron.train();
+Extract the features and labels for both training and test sets:
- multilayerPerceptron.predict(...);
+```java
+ var trainFeatures=dataProcessor.getTrainFeatures(featureColumnIndices);
+ var trainLabels=dataProcessor.getTrainLabels(targetColumnIndex);
+ var testFeatures=dataProcessor.getTestFeatures(featureColumnIndices);
+ var testLabels=dataProcessor.getTestLabels(targetColumnIndex);
+```
+
+### Step 3: Network Configuration
+
+```java
+var networkConfiguration=new NetworkConfiguration(
+ trainFeatures[0].length, // Number of input neurons
+ List.of(128,256,512), // Number of neurons in each hidden layer
+ 3, // Number of output neurons
+ 0.01, // Learning rate
+ 300, // Number of epochs
+ ActivationFunction.LEAKY_RELU, // Activation function for hidden layers
+ ActivationFunction.SOFTMAX, // Activation function for output layer
+ LossFunction.CATEGORICAL_CROSS_ENTROPY, // Loss function
+ Initialization.XAVIER, // Weight initialization for hidden layers
+ Initialization.XAVIER // Weight initialization for output layer
+ );
+```
+
+### Step 4: Training and Evaluation
+
+```java
+MultilayerPerceptron multilayerPerceptron=new MultilayerPerceptron(
+ networkConfiguration,
+ testFeatures,
+ testLabels
+ );
+ multilayerPerceptron.train(trainFeatures,trainLabels);
+ multilayerPerceptron.evaluate(testFeatures,testLabels);
+```
+
+### Results
+
+```output
+...
+MultilayerPerceptron - Best accuracy after restoring best MLP model: 98,56%
+```
### Working examples
-You can find working examples for all algorithms in the [examples](https://github.com/Samyssmile/edux/tree/main/example/src/main/java/de/example) folder.
-In all examples the IRIS or Seaborn Pinguins datasets are used.
+You can find more fully working examples for all algorithms in
+the [examples](https://github.com/Samyssmile/edux/tree/main/example/src/main/java/de/example) folder.
-#### Iris Dataset
-The IRIS dataset is a multivariate dataset introduced by the British statistician and biologist Ronald Fisher in his 1936 paper *The use of multiple measurements in taxonomic problems*
+For examples we use the
-
-## Contributions
+* [IRIS dataset](https://archive.ics.uci.edu/ml/datasets/iris).
+* [SEABORNE PENGUINS dataset](https://seaborn.pydata.org/archive/0.11/tutorial/function_overview.html).
+## Contributions
-Contributions are warmly welcomed! If you find a bug, please create an issue with a detailed description of the problem. If you wish to suggest an improvement or fix a bug, please make a pull request. Also checkout the [Rules and Guidelines](https://github.com/Samyssmile/edux/wiki/Rules-&-Guidelines-for-New-Developers) page for more information.
+Contributions are warmly welcomed! If you find a bug, please create an issue with a detailed description of the problem.
+If you wish to suggest an improvement or fix a bug, please make a pull request. Also checkout
+the [Rules and Guidelines](https://github.com/Samyssmile/edux/wiki/Rules-&-Guidelines-for-New-Developers) page for more
+information.
diff --git a/docs/javadocs/allclasses-index.html b/docs/javadocs/allclasses-index.html
index a5ccedf..fb38206 100644
--- a/docs/javadocs/allclasses-index.html
+++ b/docs/javadocs/allclasses-index.html
@@ -57,53 +57,72 @@
RandomForest Classifier
- RandomForest is an ensemble learning method, which constructs a multitude of decision trees
- at training time and outputs the class that is the mode of the classes output by
- individual trees, or a mean prediction of the individual trees (regression).
+
The MultilayerPerceptron class represents a simple feedforward neural network, which
+ consists of input, hidden, and output layers.
The SupportVectorMachine class is an implementation of a Support Vector Machine (SVM) classifier, utilizing the one-vs-one strategy for multi-class classification.
RandomForest Classifier RandomForest is an ensemble learning method, which constructs a multitude
+ of decision trees at training time and outputs the class that is the mode of the classes output
+ by individual trees, or a mean prediction of the individual trees (regression).
The SupportVectorMachine class is an implementation of a Support Vector Machine (SVM)
+ classifier, utilizing the one-vs-one strategy for multi-class classification.
Provides a common interface for machine learning classifiers within the Edux API.
-
The Classifier interface is designed to encapsulate a variety of machine learning models, offering
- a consistent approach to training, evaluating, and utilizing classifiers. Implementations are expected to handle
- specifics related to different types of classification algorithms, such as neural networks, decision trees,
- support vector machines, etc.
+
The Classifier interface is designed to encapsulate a variety of machine learning
+ models, offering a consistent approach to training, evaluating, and utilizing classifiers.
+ Implementations are expected to handle specifics related to different types of classification
+ algorithms, such as neural networks, decision trees, support vector machines, etc.
-
Each classifier must implement methods for training the model on a dataset, evaluating its performance,
- and making predictions on new, unseen data. This design allows for interchangeability of models and promotes
- a clean separation of concerns between the data processing and model training phases.
+
Each classifier must implement methods for training the model on a dataset, evaluating its
+ performance, and making predictions on new, unseen data. This design allows for
+ interchangeability of models and promotes a clean separation of concerns between the data
+ processing and model training phases.
+
+
Typical usage involves:
-
Typical usage involves:
-
Creating an instance of a class that implements Classifier.
-
Training the classifier with known data via the train method.
-
Evaluating the classifier's performance with test data via the evaluate method.
-
Applying the trained classifier to new data to make predictions via the predict method.
+
Creating an instance of a class that implements Classifier.
+
Training the classifier with known data via the train method.
+
Evaluating the classifier's performance with test data via the evaluate method.
+
Applying the trained classifier to new data to make predictions via the predict
+ method.
Implementing classes should ensure that proper validation is performed on the input data and
- any necessary pre-processing or feature scaling is applied consistent with the model's requirements.
+ any necessary pre-processing or feature scaling is applied consistent with the model's
+ requirements.
@@ -179,27 +183,24 @@
evaluate
double[][] testTargets)
Evaluates the model's performance against the provided test inputs and targets.
- This method takes a set of test inputs and their corresponding expected targets,
- applies the model to predict the outputs for the inputs, and then compares
- the predicted outputs to the expected targets to evaluate the performance
- of the model. The nature and metric of the evaluation (e.g., accuracy, MSE, etc.)
- are dependent on the specific implementation within the method.
+
This method takes a set of test inputs and their corresponding expected targets, applies the
+ model to predict the outputs for the inputs, and then compares the predicted outputs to the
+ expected targets to evaluate the performance of the model. The nature and metric of the
+ evaluation (e.g., accuracy, MSE, etc.) are dependent on the specific implementation within the
+ method.
Parameter:
-
testInputs - 2D array of double, where each inner array represents
- a single set of input values to be evaluated by the model.
-
testTargets - 2D array of double, where each inner array represents
- the expected output or target for the corresponding set
- of inputs in testInputs.
+
testInputs - 2D array of double, where each inner array represents a single set of input
+ values to be evaluated by the model.
+
testTargets - 2D array of double, where each inner array represents the expected output or
+ target for the corresponding set of inputs in testInputs.
Gibt zurück:
-
a double value representing the performance of the model when evaluated
- against the provided test inputs and targets. The interpretation of this
- value (e.g., higher is better, lower is better, etc.) depends on the
- specific evaluation metric being used.
+
a double value representing the performance of the model when evaluated against the
+ provided test inputs and targets. The interpretation of this value (e.g., higher is better,
+ lower is better, etc.) depends on the specific evaluation metric being used.
Löst aus:
-
IllegalArgumentException - if the lengths of testInputs and
- testTargets do not match, or if
- they are empty.
+
IllegalArgumentException - if the lengths of testInputs and testTargets
+ do not match, or if they are empty.
@@ -212,8 +213,7 @@
predict
Parameter:
feature - a single set of input values to be evaluated by the model.
Gibt zurück:
-
a double array representing the predicted output values for the
- provided input values.
+
a double array representing the predicted output values for the provided input values.
The DataPostProcessor interface defines a set of methods for post-processing data. This
+ typically includes normalizing data, shuffling, handling missing values, and splitting datasets.
+ Implementations should ensure that the data is properly processed to be ready for subsequent
+ analysis or machine learning tasks.
Normalizes the dataset. This typically involves scaling the values of numeric attributes so
+ that they share a common scale, often between 0 and 1, without distorting differences in the
+ ranges of values.
+
+
Gibt zurück:
+
the DataPostProcessor instance with normalized data for method chaining
Shuffles the dataset randomly. This is usually done to ensure that the data does not carry any
+ inherent bias in the order it was collected or presented.
+
+
Gibt zurück:
+
the DataPostProcessor instance with shuffled data for method chaining
Performs imputation on missing values in a specified column using the provided imputation
+ strategy. Imputation is the process of replacing missing data with substituted values.
+
+
Parameter:
+
columnName - the name of the column to apply imputation
+
imputationStrategy - the strategy to use for imputing missing values
+
Gibt zurück:
+
the DataPostProcessor instance with imputed data for method chaining
Performs imputation on missing values in a specified column index using the provided imputation
+ strategy.
+
+
Parameter:
+
columnIndex - the index of the column to apply imputation
+
imputationStrategy - the strategy to use for imputing missing values
+
Gibt zurück:
+
the DataPostProcessor instance with imputed data for method chaining
+
+
+
+
+
+
performListWiseDeletion
+
voidperformListWiseDeletion()
+
Performs list-wise deletion on the dataset. This involves removing any rows with missing values
+ to ensure the dataset is complete. This method modifies the dataset in place and does not
+ return a value.
Splits the dataset into two separate datasets according to the specified split ratio. The split
+ ratio determines the proportion of data to be used for the first dataset (e.g., training set).
+
+
Parameter:
+
splitRatio - the ratio for splitting the dataset, where 0 Ungültige Eingabe: "<" splitRatio Ungültige Eingabe: "<" 1
+
Gibt zurück:
+
a DataProcessor instance containing the first portion of the dataset according
+ to the split ratio
Loads a dataset from the specified CSV file, processes it, and returns a DataProcessor
+ that is ready to be used for further operations such as data manipulation or analysis.
Splits the dataset into two separate datasets according to the specified split ratio. The split
+ ratio determines the proportion of data to be used for the first dataset (e.g., training set).
splitRatio - the ratio for splitting the dataset, where 0 Ungültige Eingabe: "<" splitRatio Ungültige Eingabe: "<" 1
+
Gibt zurück:
+
a DataProcessor instance containing the first portion of the dataset according
+ to the split ratio
@@ -222,9 +261,21 @@
loadDataSetFromCSV
boolean skipHead,
int[] inputColumns,
int targetColumn)
+
Beschreibung aus Schnittstelle kopiert: Dataloader
+
Loads a dataset from the specified CSV file, processes it, and returns a DataProcessor
+ that is ready to be used for further operations such as data manipulation or analysis.
Normalizes the dataset. This typically involves scaling the values of numeric attributes so
+ that they share a common scale, often between 0 and 1, without distorting differences in the
+ ranges of values.
Shuffles the dataset randomly. This is usually done to ensure that the data does not carry any
+ inherent bias in the order it was collected or presented.
Performs imputation on missing values in a specified column using the provided imputation
+ strategy. Imputation is the process of replacing missing data with substituted values.
Performs list-wise deletion on the dataset. This involves removing any rows with missing values
+ to ensure the dataset is complete. This method modifies the dataset in place and does not
+ return a value.
The Dataloader interface defines a method for loading datasets from CSV files.
+ Implementations of this interface should handle the parsing of CSV files and configuration of
+ data processing according to the provided parameters.
@@ -111,7 +114,10 @@
Methodenübersicht
boolean skipHead,
int[] inputColumns,
int targetColumn)
-
+
+
Loads a dataset from the specified CSV file, processes it, and returns a DataProcessor
+ that is ready to be used for further operations such as data manipulation or analysis.
+
@@ -134,6 +140,21 @@
loadDataSetFromCSV
boolean skipHead,
int[] inputColumns,
int targetColumn)
+
Loads a dataset from the specified CSV file, processes it, and returns a DataProcessor
+ that is ready to be used for further operations such as data manipulation or analysis.
+
+
Parameter:
+
csvFile - the CSV file to load the data from
+
csvSeparator - the character that separates values in a row in the CSV file
+
skipHead - a boolean indicating whether to skip the header row (true) or not (false)
+
inputColumns - an array of indexes indicating which columns to include as input features
+
targetColumn - the index of the column to use as the output label or target for
+ predictions
+
Gibt zurück:
+
a DataProcessor object that contains the processed data
Enumerates common activation functions used in neural networks and similar machine learning architectures.
+
Enumerates common activation functions used in neural networks and similar machine learning
+ architectures.
-
Each member of this enum represents a distinct type of activation function, a critical component in
- neural networks. Activation functions determine the output of a neural network layer for a given set of
- input, and they help normalize the output of each neuron to a specific range, usually between 1 and -1 or
- between 1 and 0.
+
Each member of this enum represents a distinct type of activation function, a critical
+ component in neural networks. Activation functions determine the output of a neural network layer
+ for a given set of input, and they help normalize the output of each neuron to a specific range,
+ usually between 1 and -1 or between 1 and 0.
-
This enum simplifies the process of selecting and utilizing an activation function. It provides an
- abstraction where the user can easily switch between different functions, making it easier to experiment
- with neural network design. Additionally, each function includes a method for calculating its derivative,
- which is essential for backpropagation in neural network training.
+
This enum simplifies the process of selecting and utilizing an activation function. It
+ provides an abstraction where the user can easily switch between different functions, making it
+ easier to experiment with neural network design. Additionally, each function includes a method
+ for calculating its derivative, which is essential for backpropagation in neural network
+ training.
+
+
Available functions include:
-
Available functions include:
-
SIGMOID: Normalizes inputs between 0 and 1, crucial for binary classification.
-
RELU: Addresses the vanishing gradient problem, allowing for faster and more effective training.
-
LEAKY_RELU: Variation of RELU, prevents "dying neurons" by allowing a small gradient when the unit is not active.
-
TANH: Normalizes inputs between -1 and 1, a scaled version of the sigmoid function.
-
SOFTMAX: Converts a vector of raw scores to a probability distribution, typically used in multi-class classification.
+
SIGMOID: Normalizes inputs between 0 and 1, crucial for binary classification.
+
RELU: Addresses the vanishing gradient problem, allowing for faster and more
+ effective training.
+
LEAKY_RELU: Variation of RELU, prevents "dying neurons" by allowing a small gradient
+ when the unit is not active.
+
TANH: Normalizes inputs between -1 and 1, a scaled version of the sigmoid function.
+
SOFTMAX: Converts a vector of raw scores to a probability distribution, typically
+ used in multi-class classification.
-
Each function overrides the calculateActivation and calculateDerivative methods, providing the
- specific implementation for the activation and its derivative based on input. These are essential for the forward
- and backward passes through the network, respectively.
+
Each function overrides the calculateActivation and calculateDerivative
+ methods, providing the specific implementation for the activation and its derivative based on
+ input. These are essential for the forward and backward passes through the network, respectively.
-
Note: The SOFTMAX function additionally overrides calculateActivation for an array input,
- facilitating its common use in output layers of neural networks for classification tasks.
+
Note: The SOFTMAX function additionally overrides calculateActivation
+ for an array input, facilitating its common use in output layers of neural networks for
+ classification tasks.
Provides the classes necessary to define various activation functions used in neural networks.
-
This package is part of the larger Edux framework for educational purposes in the realm of machine learning.
- Within this package, you will find enumerations and possibly classes that represent a variety of standard
- activation functions, such as Sigmoid, TanH, ReLU, and others. These functions are fundamental components
- in the construction of neural networks, as they dictate how signals are processed as they pass from one
- neuron (or node) to the next, essentially determining the output of each neuron.
+
This package is part of the larger Edux framework for educational purposes in the realm of
+ machine learning. Within this package, you will find enumerations and possibly classes that
+ represent a variety of standard activation functions, such as Sigmoid, TanH, ReLU, and others.
+ These functions are fundamental components in the construction of neural networks, as they
+ dictate how signals are processed as they pass from one neuron (or node) to the next, essentially
+ determining the output of each neuron.
-
Each activation function contained within this package has distinct characteristics and is useful in
- different scenarios, depending on the nature of the input data, the specific architecture of the network,
- and the learning task at hand. For instance, some functions are better suited for dealing with issues like
- the vanishing gradient problem, while others might normalize input values into a certain range to aid with
- the convergence of the learning algorithm.
+
Each activation function contained within this package has distinct characteristics and is
+ useful in different scenarios, depending on the nature of the input data, the specific
+ architecture of the network, and the learning task at hand. For instance, some functions are
+ better suited for dealing with issues like the vanishing gradient problem, while others might
+ normalize input values into a certain range to aid with the convergence of the learning
+ algorithm.
-
This package is designed to offer flexibility and ease of use for those constructing machine learning
- models, as it allows for easy switching between different activation strategies, facilitating experimentation
- and learning.
+
This package is designed to offer flexibility and ease of use for those constructing machine
+ learning models, as it allows for easy switching between different activation strategies,
+ facilitating experimentation and learning.
Implements the IImputationStrategy interface to provide an average value imputation. This
+ strategy calculates the average of the non-missing numeric values in a column and substitutes the
+ missing values with this average.
+
+
It is important to note that this strategy is only applicable to columns with numeric data.
+ Attempting to use this strategy on categorical data will result in a RuntimeException.
Performs average value imputation on the provided dataset column. Missing values are identified
+ as blank strings and are replaced by the average of the non-missing values. If the column
+ contains categorical data, a runtime exception is thrown.
Defines a strategy interface for imputing missing values within a column of data. Implementations
+ of this interface should provide a concrete imputation method that can handle various types of
+ missing data according to specific rules or algorithms.
Performs imputation on the provided column data array. Missing values within the array are
+ expected to be filled with substituted values determined by the specific imputation strategy
+ implemented.
+
+
Parameter:
+
columnData - an array of String representing the data of a single column, where
+ missing values are to be imputed.
+
Gibt zurück:
+
an array of String representing the column data after imputation has been
+ performed.
Enumerates the available imputation strategies for handling missing values in datasets. Each
+ strategy is associated with a concrete implementation of IImputationStrategy that defines
+ the specific imputation behavior.
Imputation strategy that replaces missing values with the average of the non-missing values in
+ the dataset column. This strategy is suitable for numerical data only.
Imputation strategy that replaces missing values with the most frequently occurring value
+ (mode) in the dataset column. This strategy can be used for both numerical and categorical
+ data.
Implements the IImputationStrategy interface to provide a mode value imputation. This
+ strategy finds the most frequently occurring value, or mode, in a dataset column and substitutes
+ missing values with this mode.
Performs mode value imputation on the provided dataset column. Missing values are identified as
+ blank strings and are replaced by the mode of the non-missing values. If multiple modes are
+ found, the first encountered in the dataset is used.
He initialization strategy for weights. This strategy is designed for layers with ReLU
+ activation, initializing the weights with variance scaled by the size of the previous layer,
+ aiming to reduce the vanishing gradient problem.
A Decision Tree classifier for predictive modeling.
-
The DecisionTree class is a binary tree where each node represents a decision
- on a particular feature from the input feature vector, effectively partitioning the
- input space into regions with similar output labels. The tree is built recursively
- by selecting splits that minimize the Gini impurity of the resultant partitions.
+
The DecisionTree class is a binary tree where each node represents a decision on a
+ particular feature from the input feature vector, effectively partitioning the input space into
+ regions with similar output labels. The tree is built recursively by selecting splits that
+ minimize the Gini impurity of the resultant partitions.
Features:
+
-
Supports binary classification problems.
-
Utilizes the Gini impurity to determine optimal feature splits.
-
Enables control over tree depth and complexity through various hyperparameters.
+
Supports binary classification problems.
+
Utilizes the Gini impurity to determine optimal feature splits.
+
Enables control over tree depth and complexity through various hyperparameters.
Hyperparameters include:
+
-
maxDepth: The maximum depth of the tree.
-
minSamplesSplit: The minimum number of samples required to split an internal node.
-
minSamplesLeaf: The minimum number of samples required to be at a leaf node.
-
maxLeafNodes: The maximum number of leaf nodes in the tree.
+
maxDepth: The maximum depth of the tree.
+
minSamplesSplit: The minimum number of samples required to split an internal node.
+
minSamplesLeaf: The minimum number of samples required to be at a leaf node.
+
maxLeafNodes: The maximum number of leaf nodes in the tree.
Note: This class requires a thorough validation of input data and parameters, ensuring
- they are never null, have appropriate dimensions, and adhere to any other
- prerequisites or assumptions, to guarantee robustness and avoid runtime exceptions.
+
Note: This class requires a thorough validation of input data and parameters, ensuring they
+ are never null, have appropriate dimensions, and adhere to any other prerequisites or
+ assumptions, to guarantee robustness and avoid runtime exceptions.
Siehe auch:
@@ -252,25 +255,23 @@
evaluate
Beschreibung aus Schnittstelle kopiert: Classifier
Evaluates the model's performance against the provided test inputs and targets.
- This method takes a set of test inputs and their corresponding expected targets,
- applies the model to predict the outputs for the inputs, and then compares
- the predicted outputs to the expected targets to evaluate the performance
- of the model. The nature and metric of the evaluation (e.g., accuracy, MSE, etc.)
- are dependent on the specific implementation within the method.
+
This method takes a set of test inputs and their corresponding expected targets, applies the
+ model to predict the outputs for the inputs, and then compares the predicted outputs to the
+ expected targets to evaluate the performance of the model. The nature and metric of the
+ evaluation (e.g., accuracy, MSE, etc.) are dependent on the specific implementation within the
+ method.
testInputs - 2D array of double, where each inner array represents
- a single set of input values to be evaluated by the model.
-
testTargets - 2D array of double, where each inner array represents
- the expected output or target for the corresponding set
- of inputs in testInputs.
+
testInputs - 2D array of double, where each inner array represents a single set of input
+ values to be evaluated by the model.
+
testTargets - 2D array of double, where each inner array represents the expected output or
+ target for the corresponding set of inputs in testInputs.
Gibt zurück:
-
a double value representing the performance of the model when evaluated
- against the provided test inputs and targets. The interpretation of this
- value (e.g., higher is better, lower is better, etc.) depends on the
- specific evaluation metric being used.
+
a double value representing the performance of the model when evaluated against the
+ provided test inputs and targets. The interpretation of this value (e.g., higher is better,
+ lower is better, etc.) depends on the specific evaluation metric being used.
@@ -286,8 +287,7 @@
predict
Parameter:
feature - a single set of input values to be evaluated by the model.
Gibt zurück:
-
a double array representing the predicted output values for the
- provided input values.
+
a double array representing the predicted output values for the provided input values.
The KnnClassifier class provides an implementation of the k-Nearest Neighbors algorithm for classification tasks.
- It stores the training dataset and predicts the label for new data points based on the majority label of its k-nearest neighbors in the feature space.
- Distance between data points is computed using the Euclidean distance metric.
- Optionally, predictions can be weighted by the inverse of the distance to give closer neighbors higher influence.
+
The KnnClassifier class provides an implementation of the k-Nearest Neighbors algorithm
+ for classification tasks. It stores the training dataset and predicts the label for new data
+ points based on the majority label of its k-nearest neighbors in the feature space. Distance
+ between data points is computed using the Euclidean distance metric. Optionally, predictions can
+ be weighted by the inverse of the distance to give closer neighbors higher influence.
+
+
Example usage:
-
Example usage:
int k = 3; // Specify the number of neighbors to consider
KnnClassifier knn = new KnnClassifier(k);
@@ -111,7 +113,7 @@
Beschreibung aus Schnittstelle kopiert: Classifier
Evaluates the model's performance against the provided test inputs and targets.
- This method takes a set of test inputs and their corresponding expected targets,
- applies the model to predict the outputs for the inputs, and then compares
- the predicted outputs to the expected targets to evaluate the performance
- of the model. The nature and metric of the evaluation (e.g., accuracy, MSE, etc.)
- are dependent on the specific implementation within the method.
+
This method takes a set of test inputs and their corresponding expected targets, applies the
+ model to predict the outputs for the inputs, and then compares the predicted outputs to the
+ expected targets to evaluate the performance of the model. The nature and metric of the
+ evaluation (e.g., accuracy, MSE, etc.) are dependent on the specific implementation within the
+ method.
testInputs - 2D array of double, where each inner array represents
- a single set of input values to be evaluated by the model.
-
testTargets - 2D array of double, where each inner array represents
- the expected output or target for the corresponding set
- of inputs in testInputs.
+
testInputs - 2D array of double, where each inner array represents a single set of input
+ values to be evaluated by the model.
+
testTargets - 2D array of double, where each inner array represents the expected output or
+ target for the corresponding set of inputs in testInputs.
Gibt zurück:
-
a double value representing the performance of the model when evaluated
- against the provided test inputs and targets. The interpretation of this
- value (e.g., higher is better, lower is better, etc.) depends on the
- specific evaluation metric being used.
+
a double value representing the performance of the model when evaluated against the
+ provided test inputs and targets. The interpretation of this value (e.g., higher is better,
+ lower is better, etc.) depends on the specific evaluation metric being used.
@@ -256,8 +257,7 @@
predict
Parameter:
feature - a single set of input values to be evaluated by the model.
Gibt zurück:
-
a double array representing the predicted output values for the
- provided input values.
+
a double array representing the predicted output values for the provided input values.
public class MultilayerPerceptronextends Object
implements Classifier
-
The MultilayerPerceptron class represents a simple feedforward neural network,
- which consists of input, hidden, and output layers. It implements the Classifier
- interface, facilitating both the training and prediction processes on a given dataset.
+
The MultilayerPerceptron class represents a simple feedforward neural network, which
+ consists of input, hidden, and output layers. It implements the Classifier interface,
+ facilitating both the training and prediction processes on a given dataset.
-
This implementation utilizes a backpropagation algorithm for training the neural network
- to adjust weights and biases, considering a set configuration defined by NetworkConfiguration.
- The network's architecture is multi-layered, comprising one or more hidden layers in addition
- to the input and output layers. Neurons within these layers utilize activation functions defined
- per layer through the configuration.
+
This implementation utilizes a backpropagation algorithm for training the neural network to
+ adjust weights and biases, considering a set configuration defined by NetworkConfiguration. The network's architecture is multi-layered, comprising one or more hidden
+ layers in addition to the input and output layers. Neurons within these layers utilize activation
+ functions defined per layer through the configuration.
-
The training process adjusts the weights and biases of neurons within the network based on
- the error between predicted and expected outputs. Additionally, the implementation provides functionality
- to save and restore the best model achieved during training based on accuracy. Early stopping is applied
- during training to prevent overfitting and unnecessary computational expense by monitoring the performance
- improvement across epochs.
+
The training process adjusts the weights and biases of neurons within the network based on the
+ error between predicted and expected outputs. Additionally, the implementation provides
+ functionality to save and restore the best model achieved during training based on accuracy.
+ Early stopping is applied during training to prevent overfitting and unnecessary computational
+ expense by monitoring the performance improvement across epochs.
+
+
Note: This implementation logs informative messages, such as accuracy per epoch, using SLF4J logging.
+
Note: This implementation logs informative messages, such as accuracy per epoch, using SLF4J
+ logging.
Siehe auch:
@@ -242,25 +243,23 @@
evaluate
Beschreibung aus Schnittstelle kopiert: Classifier
Evaluates the model's performance against the provided test inputs and targets.
- This method takes a set of test inputs and their corresponding expected targets,
- applies the model to predict the outputs for the inputs, and then compares
- the predicted outputs to the expected targets to evaluate the performance
- of the model. The nature and metric of the evaluation (e.g., accuracy, MSE, etc.)
- are dependent on the specific implementation within the method.
+
This method takes a set of test inputs and their corresponding expected targets, applies the
+ model to predict the outputs for the inputs, and then compares the predicted outputs to the
+ expected targets to evaluate the performance of the model. The nature and metric of the
+ evaluation (e.g., accuracy, MSE, etc.) are dependent on the specific implementation within the
+ method.
testInputs - 2D array of double, where each inner array represents
- a single set of input values to be evaluated by the model.
-
testTargets - 2D array of double, where each inner array represents
- the expected output or target for the corresponding set
- of inputs in testInputs.
+
testInputs - 2D array of double, where each inner array represents a single set of input
+ values to be evaluated by the model.
+
testTargets - 2D array of double, where each inner array represents the expected output or
+ target for the corresponding set of inputs in testInputs.
Gibt zurück:
-
a double value representing the performance of the model when evaluated
- against the provided test inputs and targets. The interpretation of this
- value (e.g., higher is better, lower is better, etc.) depends on the
- specific evaluation metric being used.
+
a double value representing the performance of the model when evaluated against the
+ provided test inputs and targets. The interpretation of this value (e.g., higher is better,
+ lower is better, etc.) depends on the specific evaluation metric being used.
@@ -276,8 +275,7 @@
predict
Parameter:
input - a single set of input values to be evaluated by the model.
Gibt zurück:
-
a double array representing the predicted output values for the
- provided input values.
+
a double array representing the predicted output values for the provided input values.
RandomForest Classifier
- RandomForest is an ensemble learning method, which constructs a multitude of decision trees
- at training time and outputs the class that is the mode of the classes output by
- individual trees, or a mean prediction of the individual trees (regression).
-
- Note: Training and prediction are performed in a parallel manner using thread pooling.
+
RandomForest Classifier RandomForest is an ensemble learning method, which constructs a multitude
+ of decision trees at training time and outputs the class that is the mode of the classes output
+ by individual trees, or a mean prediction of the individual trees (regression).
+
+
Note: Training and prediction are performed in a parallel manner using thread pooling.
RandomForest handles the training of individual decision trees and their predictions, and
- determines the final prediction by voting (classification) or averaging (regression) the
- outputs of all the decision trees in the forest. RandomForest is particularly well suited
- for multiclass classification and regression on datasets with complex structures.
-
- Usage example:
-
-
+ determines the final prediction by voting (classification) or averaging (regression) the outputs
+ of all the decision trees in the forest. RandomForest is particularly well suited for multiclass
+ classification and regression on datasets with complex structures.
+
+
- Thread Safety: This class uses concurrent features but may not be entirely thread-safe
+
+
+
Thread Safety: This class uses concurrent features but may not be entirely thread-safe
and should be used with caution in a multithreaded environment.
-
feature - a single set of input values to be evaluated by the model.
Gibt zurück:
-
a double array representing the predicted output values for the
- provided input values.
+
a double array representing the predicted output values for the provided input values.
@@ -252,25 +249,23 @@
evaluate
Beschreibung aus Schnittstelle kopiert: Classifier
Evaluates the model's performance against the provided test inputs and targets.
- This method takes a set of test inputs and their corresponding expected targets,
- applies the model to predict the outputs for the inputs, and then compares
- the predicted outputs to the expected targets to evaluate the performance
- of the model. The nature and metric of the evaluation (e.g., accuracy, MSE, etc.)
- are dependent on the specific implementation within the method.
+
This method takes a set of test inputs and their corresponding expected targets, applies the
+ model to predict the outputs for the inputs, and then compares the predicted outputs to the
+ expected targets to evaluate the performance of the model. The nature and metric of the
+ evaluation (e.g., accuracy, MSE, etc.) are dependent on the specific implementation within the
+ method.
features - 2D array of double, where each inner array represents
- a single set of input values to be evaluated by the model.
-
labels - 2D array of double, where each inner array represents
- the expected output or target for the corresponding set
- of inputs in testInputs.
+
features - 2D array of double, where each inner array represents a single set of input
+ values to be evaluated by the model.
+
labels - 2D array of double, where each inner array represents the expected output or
+ target for the corresponding set of inputs in testInputs.
Gibt zurück:
-
a double value representing the performance of the model when evaluated
- against the provided test inputs and targets. The interpretation of this
- value (e.g., higher is better, lower is better, etc.) depends on the
- specific evaluation metric being used.
+
a double value representing the performance of the model when evaluated against the
+ provided test inputs and targets. The interpretation of this value (e.g., higher is better,
+ lower is better, etc.) depends on the specific evaluation metric being used.
RandomForest Classifier
- RandomForest is an ensemble learning method, which constructs a multitude of decision trees
- at training time and outputs the class that is the mode of the classes output by
- individual trees, or a mean prediction of the individual trees (regression).
+
RandomForest Classifier RandomForest is an ensemble learning method, which constructs a multitude
+ of decision trees at training time and outputs the class that is the mode of the classes output
+ by individual trees, or a mean prediction of the individual trees (regression).
public class SupportVectorMachineextends Object
implements Classifier
-
The SupportVectorMachine class is an implementation of a Support Vector Machine (SVM) classifier, utilizing the one-vs-one strategy for multi-class classification.
- This SVM implementation accepts a kernel function and trains separate binary classifiers for each pair of classes in the training set, using provided kernel function and regularization parameter C.
- During the prediction, each model in the pair casts a vote and the final predicted class is the one that gets the most votes among all binary classifiers.
+
The SupportVectorMachine class is an implementation of a Support Vector Machine (SVM)
+ classifier, utilizing the one-vs-one strategy for multi-class classification. This SVM
+ implementation accepts a kernel function and trains separate binary classifiers for each pair of
+ classes in the training set, using provided kernel function and regularization parameter C.
+ During the prediction, each model in the pair casts a vote and the final predicted class is the
+ one that gets the most votes among all binary classifiers.
+
+
Example usage:
-
Example usage:
SVMKernel kernel = ... ; // Define an appropriate SVM kernel function
double c = ... ; // Define an appropriate regularization parameter
@@ -112,7 +116,8 @@
Constructs a new instance of SupportVectorMachine with a specified kernel and regularization
+ parameter.
+
@@ -188,6 +196,30 @@
Konstruktordetails
SupportVectorMachine
publicSupportVectorMachine(SVMKernel kernel,
double c)
+
Constructs a new instance of SupportVectorMachine with a specified kernel and regularization
+ parameter.
+
+
This constructor initializes a new Support Vector Machine (SVM) for classification tasks.
+ The SVM employs a one-vs-one strategy for multi-class classification. Each model pair within
+ the SVM is trained using the provided kernel function and the regularization parameter C.
+
+
The kernel is crucial for handling non-linearly separable data by defining a new space in
+ which data points are projected. The correct choice of a kernel significantly impacts the
+ performance of the SVM. The regularization parameter C controls the trade-off between achieving
+ a low training error and a low testing error that is the ability of the SVM to generalize to
+ unseen data.
+
+
Parameter:
+
kernel - The kernel to be used for the transformation of the input space. This is
+ necessary for achieving an optimal separation in a higher-dimensional space when data is
+ not linearly separable in the original space. The kernel defines how data points in space
+ are interpreted based on their similarity.
+
c - The regularization parameter that controls the trade-off between allowing training
+ errors and enforcing rigid margins. It helps to prevent overfitting by controlling the
+ strength of the penalty for errors. A higher value of C tries to minimize the
+ classification error, potentially at the expense of simplicity, while a lower value of C
+ prioritizes simplicity, potentially allowing some misclassifications.
+
@@ -228,8 +260,7 @@
predict
Parameter:
features - a single set of input values to be evaluated by the model.
Gibt zurück:
-
a double array representing the predicted output values for the
- provided input values.
+
a double array representing the predicted output values for the provided input values.
@@ -241,25 +272,23 @@
evaluate
Beschreibung aus Schnittstelle kopiert: Classifier
Evaluates the model's performance against the provided test inputs and targets.
- This method takes a set of test inputs and their corresponding expected targets,
- applies the model to predict the outputs for the inputs, and then compares
- the predicted outputs to the expected targets to evaluate the performance
- of the model. The nature and metric of the evaluation (e.g., accuracy, MSE, etc.)
- are dependent on the specific implementation within the method.
+
This method takes a set of test inputs and their corresponding expected targets, applies the
+ model to predict the outputs for the inputs, and then compares the predicted outputs to the
+ expected targets to evaluate the performance of the model. The nature and metric of the
+ evaluation (e.g., accuracy, MSE, etc.) are dependent on the specific implementation within the
+ method.
features - 2D array of double, where each inner array represents
- a single set of input values to be evaluated by the model.
-
labels - 2D array of double, where each inner array represents
- the expected output or target for the corresponding set
- of inputs in testInputs.
+
features - 2D array of double, where each inner array represents a single set of input
+ values to be evaluated by the model.
+
labels - 2D array of double, where each inner array represents the expected output or
+ target for the corresponding set of inputs in testInputs.
Gibt zurück:
-
a double value representing the performance of the model when evaluated
- against the provided test inputs and targets. The interpretation of this
- value (e.g., higher is better, lower is better, etc.) depends on the
- specific evaluation metric being used.
+
a double value representing the performance of the model when evaluated against the
+ provided test inputs and targets. The interpretation of this value (e.g., higher is better,
+ lower is better, etc.) depends on the specific evaluation metric being used.
The SupportVectorMachine class is an implementation of a Support Vector Machine (SVM) classifier, utilizing the one-vs-one strategy for multi-class classification.
+
The SupportVectorMachine class is an implementation of a Support Vector Machine (SVM)
+ classifier, utilizing the one-vs-one strategy for multi-class classification.
Loads a dataset from the specified CSV file, processes it, and returns a DataProcessor
+ that is ready to be used for further operations such as data manipulation or analysis.
RandomForest Classifier
- RandomForest is an ensemble learning method, which constructs a multitude of decision trees
- at training time and outputs the class that is the mode of the classes output by
- individual trees, or a mean prediction of the individual trees (regression).
+
RandomForest Classifier RandomForest is an ensemble learning method, which constructs a multitude
+ of decision trees at training time and outputs the class that is the mode of the classes output
+ by individual trees, or a mean prediction of the individual trees (regression).
The SupportVectorMachine class is an implementation of a Support Vector Machine (SVM) classifier, utilizing the one-vs-one strategy for multi-class classification.
+
The SupportVectorMachine class is an implementation of a Support Vector Machine (SVM)
+ classifier, utilizing the one-vs-one strategy for multi-class classification.
diff --git a/docs/javadocs/package-search-index.js b/docs/javadocs/package-search-index.js
index 3e0dddf..a3edc4a 100644
--- a/docs/javadocs/package-search-index.js
+++ b/docs/javadocs/package-search-index.js
@@ -1 +1 @@
-packageSearchIndex = [{"l":"Alle Packages","u":"allpackages-index.html"},{"l":"de.edux.api"},{"l":"de.edux.data.provider"},{"l":"de.edux.data.reader"},{"l":"de.edux.functions.activation"},{"l":"de.edux.functions.imputation"},{"l":"de.edux.functions.initialization"},{"l":"de.edux.functions.loss"},{"l":"de.edux.math"},{"l":"de.edux.math.entity"},{"l":"de.edux.ml.decisiontree"},{"l":"de.edux.ml.knn"},{"l":"de.edux.ml.nn.config"},{"l":"de.edux.ml.nn.network"},{"l":"de.edux.ml.nn.network.api"},{"l":"de.edux.ml.randomforest"},{"l":"de.edux.ml.svm"},{"l":"de.edux.util"},{"l":"de.edux.util.math"}];updateSearchResults();
\ No newline at end of file
+packageSearchIndex = [{"l":"Alle Packages","u":"allpackages-index.html"},{"l":"de.edux.api"},{"l":"de.edux.data.provider"},{"l":"de.edux.data.reader"},{"l":"de.edux.functions.activation"},{"l":"de.edux.functions.imputation"},{"l":"de.edux.functions.initialization"},{"l":"de.edux.functions.loss"},{"l":"de.edux.math"},{"l":"de.edux.ml.decisiontree"},{"l":"de.edux.ml.knn"},{"l":"de.edux.ml.nn.config"},{"l":"de.edux.ml.nn.network"},{"l":"de.edux.ml.nn.network.api"},{"l":"de.edux.ml.randomforest"},{"l":"de.edux.ml.svm"},{"l":"de.edux.util"},{"l":"de.edux.util.math"}];updateSearchResults();
\ No newline at end of file
diff --git a/docs/javadocs/type-search-index.js b/docs/javadocs/type-search-index.js
index 6f2ed86..1adb5ad 100644
--- a/docs/javadocs/type-search-index.js
+++ b/docs/javadocs/type-search-index.js
@@ -1 +1 @@
-typeSearchIndex = [{"p":"de.edux.functions.activation","l":"ActivationFunction"},{"l":"Alle Klassen und Schnittstellen","u":"allclasses-index.html"},{"p":"de.edux.api","l":"Classifier"},{"p":"de.edux.util.math","l":"ConcurrentMatrixMultiplication"},{"p":"de.edux.data.reader","l":"CSVIDataReader"},{"p":"de.edux.data.provider","l":"DataloaderV2"},{"p":"de.edux.data.provider","l":"DataNormalizer"},{"p":"de.edux.data.provider","l":"DataPostProcessor"},{"p":"de.edux.data.provider","l":"DataProcessor"},{"p":"de.edux.ml.nn.network.api","l":"Dataset"},{"p":"de.edux.data.provider","l":"Dataset"},{"p":"de.edux.ml.decisiontree","l":"DecisionTree"},{"p":"de.edux.math","l":"Entity"},{"p":"de.edux.data.reader","l":"IDataReader"},{"p":"de.edux.ml.decisiontree","l":"IDecisionTree"},{"p":"de.edux.functions.imputation","l":"ImputationStrategy"},{"p":"de.edux.util.math","l":"IncompatibleDimensionsException"},{"p":"de.edux.ml.nn.network.api","l":"INeuron"},{"p":"de.edux.functions.initialization","l":"Initialization"},{"p":"de.edux.ml.nn.network.api","l":"IPerceptron"},{"p":"de.edux.ml.svm","l":"ISupportVectorMachine"},{"p":"de.edux.ml.knn","l":"KnnClassifier"},{"p":"de.edux.util","l":"LabelDimensionConverter"},{"p":"de.edux.functions.loss","l":"LossFunction"},{"p":"de.edux.util.math","l":"MathMatrix"},{"p":"de.edux.math","l":"MathUtil"},{"p":"de.edux.math.entity","l":"Matrix"},{"p":"de.edux.math.entity","l":"Matrix.MatrixIterator"},{"p":"de.edux.util.math","l":"MatrixOperations"},{"p":"de.edux.ml.nn.network","l":"MultilayerPerceptron"},{"p":"de.edux.ml.nn.config","l":"NetworkConfiguration"},{"p":"de.edux.data.provider","l":"Normalizer"},{"p":"de.edux.ml.randomforest","l":"RandomForest"},{"p":"de.edux.ml.randomforest","l":"Sample"},{"p":"de.edux.ml.svm","l":"SupportVectorMachine"},{"p":"de.edux.ml.svm","l":"SVMKernel"},{"p":"de.edux.ml.svm","l":"SVMModel"},{"p":"de.edux.math","l":"Validations"},{"p":"de.edux.math.entity","l":"Vector"},{"p":"de.edux.math.entity","l":"Vector.VectorIterator"}];updateSearchResults();
\ No newline at end of file
+typeSearchIndex = [{"p":"de.edux.functions.activation","l":"ActivationFunction"},{"l":"Alle Klassen und Schnittstellen","u":"allclasses-index.html"},{"p":"de.edux.functions.imputation","l":"AverageImputation"},{"p":"de.edux.api","l":"Classifier"},{"p":"de.edux.util.math","l":"ConcurrentMatrixMultiplication"},{"p":"de.edux.data.reader","l":"CSVIDataReader"},{"p":"de.edux.data.provider","l":"Dataloader"},{"p":"de.edux.data.provider","l":"DataNormalizer"},{"p":"de.edux.data.provider","l":"DataPostProcessor"},{"p":"de.edux.data.provider","l":"DataProcessor"},{"p":"de.edux.ml.nn.network.api","l":"Dataset"},{"p":"de.edux.data.provider","l":"Dataset"},{"p":"de.edux.ml.decisiontree","l":"DecisionTree"},{"p":"de.edux.math","l":"Entity"},{"p":"de.edux.data.reader","l":"IDataReader"},{"p":"de.edux.ml.decisiontree","l":"IDecisionTree"},{"p":"de.edux.functions.imputation","l":"IImputationStrategy"},{"p":"de.edux.functions.imputation","l":"ImputationStrategy"},{"p":"de.edux.util.math","l":"IncompatibleDimensionsException"},{"p":"de.edux.ml.nn.network.api","l":"INeuron"},{"p":"de.edux.functions.initialization","l":"Initialization"},{"p":"de.edux.ml.nn.network.api","l":"IPerceptron"},{"p":"de.edux.ml.svm","l":"ISupportVectorMachine"},{"p":"de.edux.ml.knn","l":"KnnClassifier"},{"p":"de.edux.util","l":"LabelDimensionConverter"},{"p":"de.edux.functions.loss","l":"LossFunction"},{"p":"de.edux.util.math","l":"MathMatrix"},{"p":"de.edux.math","l":"MathUtil"},{"p":"de.edux.math","l":"Matrix"},{"p":"de.edux.math","l":"Matrix.MatrixIterator"},{"p":"de.edux.util.math","l":"MatrixOperations"},{"p":"de.edux.functions.imputation","l":"ModeImputation"},{"p":"de.edux.ml.nn.network","l":"MultilayerPerceptron"},{"p":"de.edux.ml.nn.config","l":"NetworkConfiguration"},{"p":"de.edux.data.provider","l":"Normalizer"},{"p":"de.edux.ml.randomforest","l":"RandomForest"},{"p":"de.edux.ml.randomforest","l":"Sample"},{"p":"de.edux.ml.svm","l":"SupportVectorMachine"},{"p":"de.edux.ml.svm","l":"SVMKernel"},{"p":"de.edux.ml.svm","l":"SVMModel"},{"p":"de.edux.math","l":"Validations"},{"p":"de.edux.math","l":"Vector"},{"p":"de.edux.math","l":"Vector.VectorIterator"}];updateSearchResults();
\ No newline at end of file
diff --git a/example/build.gradle b/example/build.gradle
index c0e9d52..2a7b346 100644
--- a/example/build.gradle
+++ b/example/build.gradle
@@ -3,7 +3,7 @@ plugins {
}
group = 'io.github.samyssmile'
-version = '1.0.5'
+version = '1.0.6'
java {
toolchain {
diff --git a/example/src/main/java/de/example/nn/MultilayerNeuralNetworkExampleOnIrisDataset.java b/example/src/main/java/de/example/nn/MultilayerNeuralNetworkExampleOnIrisDataset.java
index 6384f1c..82a5a34 100644
--- a/example/src/main/java/de/example/nn/MultilayerNeuralNetworkExampleOnIrisDataset.java
+++ b/example/src/main/java/de/example/nn/MultilayerNeuralNetworkExampleOnIrisDataset.java
@@ -50,7 +50,7 @@ public static void main(String[] args) {
// - 2 Hidden Layer with 12 and 6 Neurons
// - 3 Output Neurons
// - Learning Rate of 0.1
- // - 1000 Epochs
+ // - 300 Epochs
// - Leaky ReLU as Activation Function for Hidden Layers
// - Softmax as Activation Function for Output Layer
// - Categorical Cross Entropy as Loss Function
@@ -61,7 +61,7 @@ public static void main(String[] args) {
trainFeatures[0].length,
List.of(128, 256, 512),
3,
- 0.01,
+ 0.005,
300,
ActivationFunction.LEAKY_RELU,
ActivationFunction.SOFTMAX,
diff --git a/lib/build.gradle b/lib/build.gradle
index efd23df..4fe088a 100644
--- a/lib/build.gradle
+++ b/lib/build.gradle
@@ -5,7 +5,7 @@ plugins {
}
group = 'io.github.samyssmile'
-version = '1.0.5'
+version = '1.0.6'
java {
toolchain {
@@ -58,7 +58,7 @@ publishing {
mavenJava(MavenPublication) {
groupId = 'io.github.samyssmile'
artifactId = 'edux'
- version = '1.0.5'
+ version = '1.0.6'
from components.java
artifact sourceJar
artifact javadocJar
diff --git a/lib/src/main/java/de/edux/data/provider/DataPostProcessor.java b/lib/src/main/java/de/edux/data/provider/DataPostProcessor.java
index 732540a..e0330d8 100644
--- a/lib/src/main/java/de/edux/data/provider/DataPostProcessor.java
+++ b/lib/src/main/java/de/edux/data/provider/DataPostProcessor.java
@@ -3,18 +3,74 @@
import de.edux.functions.imputation.ImputationStrategy;
import java.util.List;
+/**
+ * The {@code DataPostProcessor} interface defines a set of methods for post-processing data. This
+ * typically includes normalizing data, shuffling, handling missing values, and splitting datasets.
+ * Implementations should ensure that the data is properly processed to be ready for subsequent
+ * analysis or machine learning tasks.
+ */
public interface DataPostProcessor {
+
+ /**
+ * Normalizes the dataset. This typically involves scaling the values of numeric attributes so
+ * that they share a common scale, often between 0 and 1, without distorting differences in the
+ * ranges of values.
+ *
+ * @return the {@code DataPostProcessor} instance with normalized data for method chaining
+ */
DataPostProcessor normalize();
+ /**
+ * Shuffles the dataset randomly. This is usually done to ensure that the data does not carry any
+ * inherent bias in the order it was collected or presented.
+ *
+ * @return the {@code DataPostProcessor} instance with shuffled data for method chaining
+ */
DataPostProcessor shuffle();
+ /**
+ * Performs imputation on missing values in a specified column using the provided imputation
+ * strategy. Imputation is the process of replacing missing data with substituted values.
+ *
+ * @param columnName the name of the column to apply imputation
+ * @param imputationStrategy the strategy to use for imputing missing values
+ * @return the {@code DataPostProcessor} instance with imputed data for method chaining
+ */
DataPostProcessor imputation(String columnName, ImputationStrategy imputationStrategy);
+ /**
+ * Performs imputation on missing values in a specified column index using the provided imputation
+ * strategy.
+ *
+ * @param columnIndex the index of the column to apply imputation
+ * @param imputationStrategy the strategy to use for imputing missing values
+ * @return the {@code DataPostProcessor} instance with imputed data for method chaining
+ */
DataPostProcessor imputation(int columnIndex, ImputationStrategy imputationStrategy);
+ /**
+ * Performs list-wise deletion on the dataset. This involves removing any rows with missing values
+ * to ensure the dataset is complete. This method modifies the dataset in place and does not
+ * return a value.
+ */
void performListWiseDeletion();
+ /**
+ * Retrieves the processed dataset as a list of string arrays. Each string array represents a row
+ * in the dataset.
+ *
+ * @return a list of string arrays representing the dataset
+ */
List getDataset();
+ /**
+ * Splits the dataset into two separate datasets according to the specified split ratio. The split
+ * ratio determines the proportion of data to be used for the first dataset (e.g., training set).
+ *
+ * @param splitRatio the ratio for splitting the dataset, where 0 < splitRatio < 1
+ * @return a {@code DataProcessor} instance containing the first portion of the dataset according
+ * to the split ratio
+ * @throws IllegalArgumentException if the split ratio is not between 0 and 1
+ */
DataProcessor split(double splitRatio);
}
diff --git a/lib/src/main/java/de/edux/data/provider/DataProcessor.java b/lib/src/main/java/de/edux/data/provider/DataProcessor.java
index 4becc52..ad36ef7 100644
--- a/lib/src/main/java/de/edux/data/provider/DataProcessor.java
+++ b/lib/src/main/java/de/edux/data/provider/DataProcessor.java
@@ -7,7 +7,7 @@
import org.slf4j.Logger;
import org.slf4j.LoggerFactory;
-public class DataProcessor implements DataPostProcessor, Dataset, DataloaderV2 {
+public class DataProcessor implements DataPostProcessor, Dataset, Dataloader {
private static final Logger LOG = LoggerFactory.getLogger(DataProcessor.class);
private final IDataReader dataReader;
private final Normalizer normalizer;
diff --git a/lib/src/main/java/de/edux/data/provider/Dataloader.java b/lib/src/main/java/de/edux/data/provider/Dataloader.java
new file mode 100644
index 0000000..f905a8c
--- /dev/null
+++ b/lib/src/main/java/de/edux/data/provider/Dataloader.java
@@ -0,0 +1,26 @@
+package de.edux.data.provider;
+
+import java.io.File;
+
+/**
+ * The {@code Dataloader} interface defines a method for loading datasets from CSV files.
+ * Implementations of this interface should handle the parsing of CSV files and configuration of
+ * data processing according to the provided parameters.
+ */
+public interface Dataloader {
+ /**
+ * Loads a dataset from the specified CSV file, processes it, and returns a {@code DataProcessor}
+ * that is ready to be used for further operations such as data manipulation or analysis.
+ *
+ * @param csvFile the CSV file to load the data from
+ * @param csvSeparator the character that separates values in a row in the CSV file
+ * @param skipHead a boolean indicating whether to skip the header row (true) or not (false)
+ * @param inputColumns an array of indexes indicating which columns to include as input features
+ * @param targetColumn the index of the column to use as the output label or target for
+ * predictions
+ * @return a {@code DataProcessor} object that contains the processed data
+ * @throws IllegalArgumentException if parameters are invalid or if the file cannot be processed
+ */
+ DataProcessor loadDataSetFromCSV(
+ File csvFile, char csvSeparator, boolean skipHead, int[] inputColumns, int targetColumn);
+}
diff --git a/lib/src/main/java/de/edux/data/provider/DataloaderV2.java b/lib/src/main/java/de/edux/data/provider/DataloaderV2.java
deleted file mode 100644
index f7ff2d1..0000000
--- a/lib/src/main/java/de/edux/data/provider/DataloaderV2.java
+++ /dev/null
@@ -1,8 +0,0 @@
-package de.edux.data.provider;
-
-import java.io.File;
-
-public interface DataloaderV2 {
- DataProcessor loadDataSetFromCSV(
- File csvFile, char csvSeparator, boolean skipHead, int[] inputColumns, int targetColumn);
-}
diff --git a/lib/src/main/java/de/edux/functions/imputation/AverageImputation.java b/lib/src/main/java/de/edux/functions/imputation/AverageImputation.java
index 3384db4..f553faf 100644
--- a/lib/src/main/java/de/edux/functions/imputation/AverageImputation.java
+++ b/lib/src/main/java/de/edux/functions/imputation/AverageImputation.java
@@ -3,7 +3,27 @@
import java.util.Arrays;
import java.util.List;
+/**
+ * Implements the {@code IImputationStrategy} interface to provide an average value imputation. This
+ * strategy calculates the average of the non-missing numeric values in a column and substitutes the
+ * missing values with this average.
+ *
+ *
It is important to note that this strategy is only applicable to columns with numeric data.
+ * Attempting to use this strategy on categorical data will result in a {@code RuntimeException}.
+ */
public class AverageImputation implements IImputationStrategy {
+ /**
+ * Performs average value imputation on the provided dataset column. Missing values are identified
+ * as blank strings and are replaced by the average of the non-missing values. If the column
+ * contains categorical data, a runtime exception is thrown.
+ *
+ * @param datasetColumn an array of {@code String} representing the column data with potential
+ * missing values.
+ * @return an array of {@code String} where missing values have been imputed with the average of
+ * non-missing values.
+ * @throws RuntimeException if the column data contains categorical values which cannot be
+ * averaged.
+ */
@Override
public String[] performImputation(String[] datasetColumn) {
checkIfColumnContainsCategoricalValues(datasetColumn);
diff --git a/lib/src/main/java/de/edux/functions/imputation/IImputationStrategy.java b/lib/src/main/java/de/edux/functions/imputation/IImputationStrategy.java
index f500c4e..1f47549 100644
--- a/lib/src/main/java/de/edux/functions/imputation/IImputationStrategy.java
+++ b/lib/src/main/java/de/edux/functions/imputation/IImputationStrategy.java
@@ -1,5 +1,21 @@
package de.edux.functions.imputation;
+/**
+ * Defines a strategy interface for imputing missing values within a column of data. Implementations
+ * of this interface should provide a concrete imputation method that can handle various types of
+ * missing data according to specific rules or algorithms.
+ */
public interface IImputationStrategy {
+
+ /**
+ * Performs imputation on the provided column data array. Missing values within the array are
+ * expected to be filled with substituted values determined by the specific imputation strategy
+ * implemented.
+ *
+ * @param columnData an array of {@code String} representing the data of a single column, where
+ * missing values are to be imputed.
+ * @return an array of {@code String} representing the column data after imputation has been
+ * performed.
+ */
String[] performImputation(String[] columnData);
}
diff --git a/lib/src/main/java/de/edux/functions/imputation/ImputationStrategy.java b/lib/src/main/java/de/edux/functions/imputation/ImputationStrategy.java
index df799e4..7ef265b 100644
--- a/lib/src/main/java/de/edux/functions/imputation/ImputationStrategy.java
+++ b/lib/src/main/java/de/edux/functions/imputation/ImputationStrategy.java
@@ -1,15 +1,42 @@
package de.edux.functions.imputation;
+/**
+ * Enumerates the available imputation strategies for handling missing values in datasets. Each
+ * strategy is associated with a concrete implementation of {@code IImputationStrategy} that defines
+ * the specific imputation behavior.
+ */
public enum ImputationStrategy {
+ /**
+ * Imputation strategy that replaces missing values with the average of the non-missing values in
+ * the dataset column. This strategy is suitable for numerical data only.
+ */
AVERAGE(new AverageImputation()),
+
+ /**
+ * Imputation strategy that replaces missing values with the most frequently occurring value
+ * (mode) in the dataset column. This strategy can be used for both numerical and categorical
+ * data.
+ */
MODE(new ModeImputation());
private final IImputationStrategy imputation;
+ /**
+ * Constructor for the enum that associates each imputation strategy with a concrete {@code
+ * IImputationStrategy} implementation.
+ *
+ * @param imputation the imputation strategy implementation
+ */
ImputationStrategy(IImputationStrategy imputation) {
this.imputation = imputation;
}
+ /**
+ * Retrieves the {@code IImputationStrategy} implementation associated with the imputation
+ * strategy.
+ *
+ * @return the imputation strategy implementation
+ */
public IImputationStrategy getImputation() {
return this.imputation;
}
diff --git a/lib/src/main/java/de/edux/functions/imputation/ModeImputation.java b/lib/src/main/java/de/edux/functions/imputation/ModeImputation.java
index 40e5dd0..fa2b670 100644
--- a/lib/src/main/java/de/edux/functions/imputation/ModeImputation.java
+++ b/lib/src/main/java/de/edux/functions/imputation/ModeImputation.java
@@ -2,7 +2,23 @@
import java.util.*;
+/**
+ * Implements the {@code IImputationStrategy} interface to provide a mode value imputation. This
+ * strategy finds the most frequently occurring value, or mode, in a dataset column and substitutes
+ * missing values with this mode.
+ */
public class ModeImputation implements IImputationStrategy {
+
+ /**
+ * Performs mode value imputation on the provided dataset column. Missing values are identified as
+ * blank strings and are replaced by the mode of the non-missing values. If multiple modes are
+ * found, the first encountered in the dataset is used.
+ *
+ * @param datasetColumn an array of {@code String} representing the column data with potential
+ * missing values.
+ * @return an array of {@code String} where missing values have been imputed with the mode of
+ * non-missing values.
+ */
@Override
public String[] performImputation(String[] datasetColumn) {
diff --git a/lib/src/main/java/de/edux/functions/initialization/Initialization.java b/lib/src/main/java/de/edux/functions/initialization/Initialization.java
index a607ab9..35ede79 100644
--- a/lib/src/main/java/de/edux/functions/initialization/Initialization.java
+++ b/lib/src/main/java/de/edux/functions/initialization/Initialization.java
@@ -1,7 +1,22 @@
package de.edux.functions.initialization;
+/**
+ * Enumerates strategies for initializing weights in neural network layers, providing methods to
+ * apply these strategies to given weight arrays.
+ */
public enum Initialization {
+ /**
+ * Enumerates strategies for initializing weights in neural network layers, providing methods to
+ * apply these strategies to given weight arrays.
+ */
XAVIER {
+ /**
+ * Applies Xavier initialization to the provided weights array using the input size.
+ *
+ * @param inputSize the size of the layer input
+ * @param weights the array of weights to be initialized
+ * @return the initialized weights array
+ */
@Override
public double[] weightInitialization(int inputSize, double[] weights) {
double xavier = Math.sqrt(6.0 / (inputSize + 1));
@@ -11,7 +26,20 @@ public double[] weightInitialization(int inputSize, double[] weights) {
return weights;
}
},
+
+ /**
+ * He initialization strategy for weights. This strategy is designed for layers with ReLU
+ * activation, initializing the weights with variance scaled by the size of the previous layer,
+ * aiming to reduce the vanishing gradient problem.
+ */
HE {
+ /**
+ * Applies He initialization to the provided weights array using the input size.
+ *
+ * @param inputSize the size of the layer input
+ * @param weights the array of weights to be initialized
+ * @return the initialized weights array
+ */
@Override
public double[] weightInitialization(int inputSize, double[] weights) {
double he = Math.sqrt(2.0 / inputSize);
diff --git a/lib/src/main/java/de/edux/math/entity/Matrix.java b/lib/src/main/java/de/edux/math/Matrix.java
similarity index 96%
rename from lib/src/main/java/de/edux/math/entity/Matrix.java
rename to lib/src/main/java/de/edux/math/Matrix.java
index 804aa05..c8f63ae 100644
--- a/lib/src/main/java/de/edux/math/entity/Matrix.java
+++ b/lib/src/main/java/de/edux/math/Matrix.java
@@ -1,8 +1,5 @@
-package de.edux.math.entity;
+package de.edux.math;
-import de.edux.math.Entity;
-import de.edux.math.MathUtil;
-import de.edux.math.Validations;
import java.util.Iterator;
import java.util.NoSuchElementException;
diff --git a/lib/src/main/java/de/edux/math/entity/Vector.java b/lib/src/main/java/de/edux/math/Vector.java
similarity index 97%
rename from lib/src/main/java/de/edux/math/entity/Vector.java
rename to lib/src/main/java/de/edux/math/Vector.java
index e19e321..66730bf 100644
--- a/lib/src/main/java/de/edux/math/entity/Vector.java
+++ b/lib/src/main/java/de/edux/math/Vector.java
@@ -1,7 +1,5 @@
-package de.edux.math.entity;
+package de.edux.math;
-import de.edux.math.Entity;
-import de.edux.math.Validations;
import java.util.Arrays;
import java.util.Iterator;
import java.util.NoSuchElementException;
diff --git a/lib/src/main/java/de/edux/math/package-info.java b/lib/src/main/java/de/edux/math/package-info.java
new file mode 100644
index 0000000..2d30553
--- /dev/null
+++ b/lib/src/main/java/de/edux/math/package-info.java
@@ -0,0 +1,5 @@
+/**
+ * Provides the classes necessary for mathematical operations and data manipulation within the Edux
+ * framework.
+ */
+package de.edux.math;
diff --git a/lib/src/main/java/de/edux/ml/nn/config/package-info.java b/lib/src/main/java/de/edux/ml/nn/config/package-info.java
new file mode 100644
index 0000000..6bbb23d
--- /dev/null
+++ b/lib/src/main/java/de/edux/ml/nn/config/package-info.java
@@ -0,0 +1,2 @@
+/** Classes for the configuration of the neural network. */
+package de.edux.ml.nn.config;
diff --git a/lib/src/test/java/de/edux/math/entity/MatrixTest.java b/lib/src/test/java/de/edux/math/entity/MatrixTest.java
index 7f2cee3..cedfa93 100644
--- a/lib/src/test/java/de/edux/math/entity/MatrixTest.java
+++ b/lib/src/test/java/de/edux/math/entity/MatrixTest.java
@@ -2,6 +2,7 @@
import static org.junit.jupiter.api.Assertions.assertEquals;
+import de.edux.math.Matrix;
import org.junit.jupiter.api.BeforeEach;
import org.junit.jupiter.api.Test;
diff --git a/lib/src/test/java/de/edux/math/entity/VectorTest.java b/lib/src/test/java/de/edux/math/entity/VectorTest.java
index fb924f3..e7c90d5 100644
--- a/lib/src/test/java/de/edux/math/entity/VectorTest.java
+++ b/lib/src/test/java/de/edux/math/entity/VectorTest.java
@@ -2,6 +2,7 @@
import static org.junit.jupiter.api.Assertions.assertEquals;
+import de.edux.math.Vector;
import org.junit.jupiter.api.BeforeAll;
import org.junit.jupiter.api.Test;